Clinical Identifier Protection via Hybrid Entity Redaction (CIPHER)
The Scale of the Problem
Clinical notes are among the most valuable yet underutilized resources in healthcare. These documents capture the evolving narrative of patient conditions, the nuanced reasoning of providers, and critical context about treatment decisions and outcomes. The scale is immense: the U.S. healthcare system generates documentation from over one billion physician office visits [7], 34 million hospital admissions [2], and 140 million emergency department encounters [8] annually, while a typical hospital generates 50 petabytes of data per year [9].
At Latent, we're researching and developing AI systems to improve patient care: Predicting disease progression before symptoms manifest, estimating readmission risk for proactive interventions, screening populations for specialized care, and providing real-time clinical decision support. The clinical depth needed to power these applications exists in provider documentation, but only becomes accessible once PHI is reliably removed. Robust, scalable de-identification is what closes that gap.
.png)
Figure 1. Synthetically generated example for purpose of demonstration
Clinical de-identification is a particularly challenging NER problem. The task requires detecting 18 categories of entities. From obvious identifiers like names and social security numbers to subtle cases like dates (which can be clinically relevant when describing disease progression) and geographic data (which provides context about healthcare access). Identifiers appear in diverse contexts: a given name might be the patient, a family member mentioned in social history or a provider signing a note. The system must distinguish between PHI-like patterns and legitimate clinical terms: is "Dr. Smith" a provider name or a medication brand?
When optimized for the specific domain and task, large language models can identify PHI accurately.But their cost, latency, and non-determinism make them impractical at scale. Rule-based systems achieve high precision on common patterns but struggle with the linguistic variance of real clinical documents, abbreviations, misspellings, and context-dependent identifiers frequently evade detection. We needed something that combines LLM-level accuracy with the speed, cost-efficiency and determinism that healthcare deployments demand.
We leverage an ensemble of LLMs to generate probability distributions over BIO labels, then distill that knowledge into a 1.5B-parameter encoder-based token classifier, achieving >98% recall on expert-labeled evaluation sets while reducing inference cost by 1000× and latency to ~50ms per document. Our approach combines outlier-aware dataset construction that oversamples challenging cases, soft-label distillation that preserves ensemble uncertainty at ambiguous entity boundaries, and a vectorized CRF implementation for efficient structured prediction. The result is a production-ready system that combines LLM-level accuracy with the determinism and efficiency that healthcare deployments demand.

Figure 2. High-Level System Architecture
Teaching a Small Model to See PHI
Our pipeline operates in four stages: 1) stratify and select a representative subsample of our full distribution using lexical and semantic features, 2) manually label this subset to produce an expert-labeled gold set of high-quality examples, 3) use an ensemble of open-source, high-reasoning LLMs to scale this with soft labels, 4) train a compact encoder on this scaled silver set to distill the identification behavior. The key insight is that we can use expensive, high-quality LLM annotations at training time while deploying a fast, deterministic encoder at inference time, capturing the best of both paradigms.
Curating the Training Data
| Feature | Description |
|---|---|
text_length | Character count of the note |
unique_tokens | Number of unique tokens (tiktoken cl100k_base) |
type_token_ratio | Lexical diversity: unique words / total words |
compression_ratio | gzip compressed size / raw size (lower = more repetitive) |
digit_share | Fraction of characters that are digits |
longest_numeric_run | Length of longest consecutive digit sequence |
non_letter_symbol_pct | Percentage of non-alphabetic characters |
mean_corpus_similarity | Mean TF-IDF cosine similarity to other notes |
regex_counts__date | Count of date patterns (MM/DD/YYYY, YYYY-MM-DD, etc.) |
regex_counts__phone | Count of phone patterns (XXX-XXX-XXXX) |
regex_counts__ssn | Count of SSN patterns (XXX-XX-XXXX) |
regex_counts__email | Count of email patterns |
regex_counts__mrn | Count of MRN patterns ("MRN: 12345") |
regex_counts__address | Count of street address patterns |
Table 1. Note Features for Outlier Detection.
We construct training datasets from clinical notes sourced across our partner healthcare systems, ensuring our models learn from diverse documentation styles, patient populations, and clinical workflows. A random sample would over-represent high-volume note types (e.g., progress notes) while under-sampling rare but challenging categories (e.g., operative reports, discharge summaries). Rather than accepting this bias, we deliberately oversample challenging cases. We compute robust Z-scores (using median absolute deviation) across 14 lexical and semantic features including text length, lexical diversity, PHI density, formatting patterns, corpus similarity, and regex-matched identifier counts. Descriptions of these features are included in table 1. We also stratify across document-type and organization groups, flagging notes with Z > 2.0 as outliers. The final dataset uses 90% stratified random sampling and 10% outlier oversampling, balancing representative coverage with sufficient exposure to edge cases. We then deploy a K=8 ensemble of OSS LLM annotators to produce soft labels across ~15,000 clinical notes, a labeling effort that cost ~$2,500 and a couple orders of magnitude of wall-clock time.
The Teacher: A Recall-Optimized LLM Ensemble
Rather than taking the ensemble's majority vote as a hard label, we preserve the full vote distribution as a soft label. For each token at position tt, the soft label distribution is:
where \(K\) is the number of ensemble members, \(v_t^{(k)}\) is the label assigned to token \(t\) by the \(k\)-th annotator, \(\mathbb{1}[\cdot]\) is the indicator function, and \(\mathcal{L}\) is the set of BIO labels. When 6 out of 8 voters agree a token is B-NAME and 2 say O, the resulting distribution [0.75, 0.25, 0, ...] encodes genuine boundary uncertainty that hard labels would discard. This is closely related to standard knowledge distillation [4], where a student trains on a teacher's softmax outputs rather than hard labels — but rather than temperature-scaling a single model's logits, our soft targets arise naturally from ensemble disagreement. Ensemble vote distributions are better calibrated [5], allow us to follow diverse reasoning trajectories, isolate genuine model disagreement from inherent ambiguity, and surface correlated failure modes that no single reasoning path can reveal.
The Student: Encoder + CRF
Our student combines a Stella 1.5B encoder, a bidirectional transformer based on Qwen2-1.5B, converted via the LLM2Vec approach [3]. The causal attention mask is replaced with a fully bidirectional mask with a classification head and a Conditional Random Field (CRF) layer. The CRF models the joint probability of the entire label sequence rather than making independent per-token predictions:
where the score function is:
and \(\mathbf{E} \in \mathbb{R}^{T \times |\mathcal{L}|}\) are the emission scores from the classification head, and \(\mathbf{A} \in \mathbb{R}^{|\mathcal{L}| \times |\mathcal{L}|}\) is the learned transition matrix where \(A_{ij}\) represents the score for transitioning from PHI label \(i\) to label \(j\).
Unlike independent token classifiers, the CRF models the conditional probability of the entire label sequence, learning transition probabilities that capture dependencies between adjacent labels. This enforces structural constraints that per-token classification cannot — an I-NAME tag should only follow B-NAME or I-NAME, never appear after O or a different entity type. At inference, we introduce a vectorized batched Viterbi algorithm that processes all sequences in a batch simultaneously, reducing GPU-CPU synchronizations from \(O(B \cdot T)\) to \(O(T)\) and achieving 20-50× speedup over naive sequential decoding.

Figure 3. Encoder + CRF Architecture
Training Objective
Training combines two complementary losses:
The distillation term is a focal-modulated soft cross-entropy that addresses the severe class imbalance between PHI and non-PHI tokens. The vast majority of tokens are non-PHI, and unmodulated cross-entropy would let the model minimize loss by confidently predicting "O" everywhere. We apply focal modulation [6]:
With γ=2.0, a token classified with 95% confidence contributes 400× less to the loss than one at 50%, forcing the model to spend its capacity on ambiguous tokens near entity boundaries rather than easy non-PHI tokens in the middle of sentences. The CRF loss operates on hard labels derived from the soft distribution (\(\hat{\mathbf{y}} = \arg\max \mathbf{p}\)) and penalizes structurally invalid sequences:
where the numerator scores the correct label sequence and the denominator marginalizes over all possible sequences via the forward algorithm. Together, the focal distillation loss trains the model to classify individual tokens accurately while the CRF loss ensures coherent entity sequences.
What Worked (and What Didn't)
CIPHER matches LLM-level accuracy at a fraction of the cost. On our held-out set of ~195 expert-annotated clinical notes, CIPHER achieves 97.9% recall and 93.85% F1, which actually exceeds its teacher LLM(GPT-OSS-120B) on recall (97.9% vs 97.5%) and trails the best frontier model by only 0.2%. The student surpassing its teacher is a known property of distillation: the ensemble's soft labels encode richer signal than any individual member provides. Critically, inference drops from seconds per document to ~50ms, a 1000× cost reduction that makes processing these documents tractable.
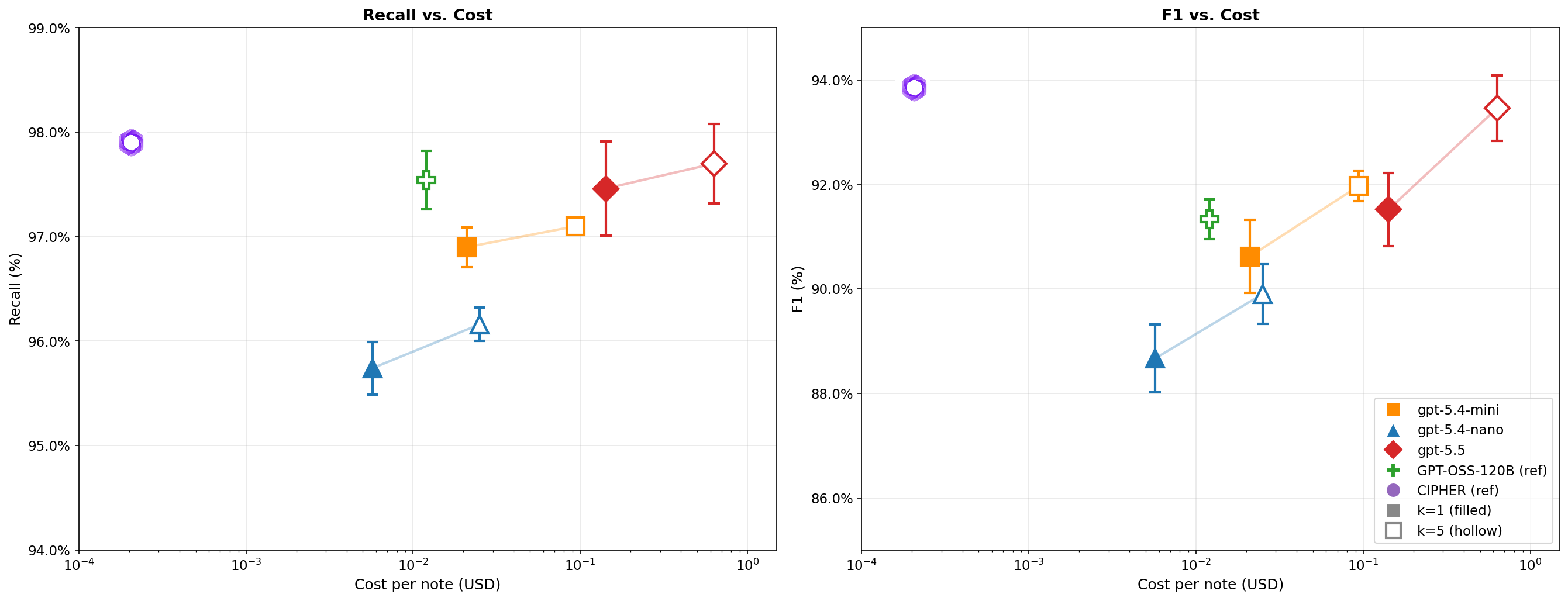
Figure 4.CIPHER vs. Decoder-Based Approaches: De-identification recall(left) and F1 (right) vs. cost per note for CIPHER and frontier LLM baselines on 195 expert-annotatedclinical notes. Frontier models (gpt-5.4-nano, gpt-5.4-mini, gpt-5.5) are shown with single-shot (k=1, filled) and 5-voter ensemble (k=5; majority vote, hollow) configurations; faded lines connect paired configurations. GPT-OSS-120B (CIPHER's teacher) is shown for reference. Error bars are ±1 SD over 10 trials. Both axes are log-scaled; cost is computed from per-token pricing (LLMs) or H100 compute time (CIPHER).
Soft labels are the single most impactful component.
| Configuration | Recall | Precision | F1 |
|---|---|---|---|
| Full System | 97.9% | 89.1% | 93.85% |
| w/o CRF | 96.65% | 89.04% | 92.69% |
| w/o Focal Loss | 96.96% | 90.53% | 93.64% |
| w/o Soft Labels | 97.43% | 84.31% | 90.40% |
Table 2.Architecture ablation
Removing soft labels and training on hard argmax labels instead drops F1 by 3.45 points — precision falls by 5% as the model, lacking information about inter-class similarity and uncertainty, treats every teacher prediction with equal confidence and aggressively labels borderline tokens as PHI. The CRF contributes +1.16% F1 through structural consistency, primarily benefiting multi-token entities like addresses and full names. Focal loss provides a modest recall boost at a small precision cost — consistent with its design intent of pushing the model toward catching more PHI. The lesson: capturing teacher uncertainty matters more than any individual architectural choice.
Scaling behavior is log-linear with diminishing returns.

Figure 5.Line plot showing soft recall, hard recall, and F1 score as a function of training set size (log scale on x-axis). Include error bars if multiple runs. Add a dashed line showing asymptotic performance to indicate when returns diminish.
Recall scales from 86.8% at 500 notes to 97.9% at 15,000, with most of the relative gain coming early with the first 2,000 notes improve recall by ~8 points. This suggests ~5,000 notes captures most of the recall ceiling, making our choice of 15,000 conservative but thorough.
Real data beats synthetic data — decisively. We evaluated mixtures of synthetic clinical notes (Nemotron-PII [10]) and authentic notes, holding total dataset size fixed at 10,000.
| Synthetic % | Authentic | Authentic Eval Recall | Authentic Eval F1 | Synthetic Eval Recall | Synthetic Eval F1 |
|---|---|---|---|---|---|
| 100% | 0 | 78.2% | 76.5% | 96.8% | 95.1% |
| 75% | 25% | 85.6% | 83.9% | 94.1% | 92.7% |
| 50% | 50% | 91.3% | 89.8% | 94.5% | 92.0% |
| 25% | 75% | 96.1% | 93.2% | 97.2% | 93.8% |
| 0% | 100% | 97.4% | 93.6% | 95.9% | 94.3% |
Table 3. Performance by data mixture
A model trained entirely on synthetic data achieves 96.8% recall on synthetic test data but only 78.2% on authentic clinical notes — a 19-point gap. Each 25% shift toward authentic data closes this gap steeply, with authentic eval recall climbing from 78.2% to 97.4%. The model trained on 100% authentic data actually achieves the highest recall on both evaluation sets, outperforming even the model trained entirely on synthetic data on its own test set. The takeaway for clinical NLP broadly: synthetic data can demonstrate that a method works in principle, but the distribution mismatch between generated and authentic text translates directly into missed PHI at inference time.
De-identification preserves clinical meaning. A critical concern with any de-identification system is whether it preserves the clinical substance of the original text. Removing identifiers provides little value if the process also strips away the context and nuance that make the original document informative.
| Metric | Value |
|---|---|
| Paired Cosine Similarity | 0.985 |
| KNN Recall (k=10) | 0.956 |
| Wasserstein Distance | 0.064 |
Table 4. Preservation of clinical semanticity
We embed each note before and after de-identification using Clinical ModernBERT [11] and measure semantic preservation along three axes. Paired cosine similarity is 0.9855, which shows that de-identification shifts the average note's embedding by less than 0.5%. KNN recall (k=10) is 0.956, meaning 95.6% of neighbor relationships survive de-identification. Wasserstein distance is 0.064, well below the 0.1 threshold, indicating no systematic distributional distortion. These results validate a core advantage of our token-level approach over generative de-identification methods [1]: because we identify and mask specific character spans without touching surrounding text, we cannot round a lab value, compress a symptom timeline, or drop a negation. Clinical meaning is preserved by construction.
What's Next
CIPHER demonstrates that knowledge distillation can transfer frontier-level PHI detection to a 1.5B-parameter encoder at 1000× lower inference cost, achieving 97.9% recall at ~50ms per document.
Our ablations show that the choice of training signal matters more than architecture. Soft labels contributed the largest improvement (+3.45 F1), encoding boundary uncertainty and inter-class similarity that hard labels discard. The CRF adds structural consistency (+1.16% F1) by enforcing valid BIO transitions, and focal loss shifts gradient signal toward rare PHI tokens at a modest precision cost. The synthetic data experiments reinforce that models need to train on the distribution they will serve — the 19-point recall gap between synthetic-only and authentic-only training is a statement about distributional fidelity, not data quantity.
Our semantic preservation metrics (0.985 cosine similarity, 0.956 KNN recall) confirm that token-level de-identification maintains clinical meaning by construction. CIPHER is one step in our broader mission of delivering personalized, provider-grade care for every patient.




